 Photo by rawpixel on Unsplash
Photo by rawpixel on UnsplashBackground
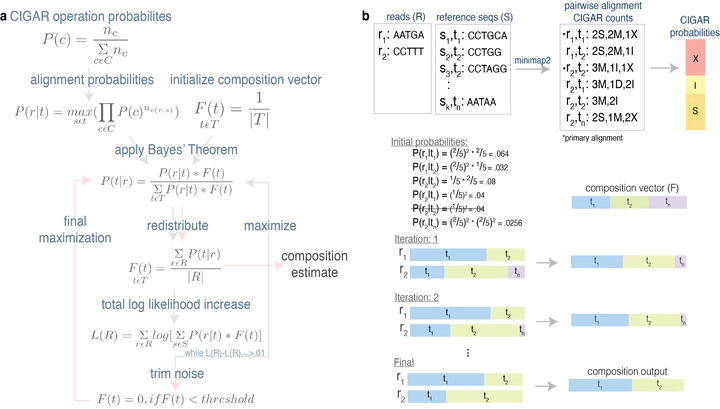
16S rRNA based analysis is the established standard for elucidating microbial community composition. While short read 16S analyses are largely confined to genus-level resolution at best since only a portion of the gene is sequenced, full-length 16S sequences have the potential to provide species-level accuracy. However, existing taxonomic identification algorithms are not optimized for the increased read length and error rate of long-read data. Here we present Emu, a novel approach that employs an expectation-maximization (EM) algorithm to generate taxonomic abundance profiles from full-length 16S rRNA reads. Results produced from one simulated data set and two mock communities prove Emu capable of accurate microbial community profiling while obtaining fewer false positives and false negatives than alternative methods. Additionally, we illustrate a real-world application of our new software by comparing clinical sample composition estimates generated by an established whole-genome shotgun sequencing workflow to those returned by full-length 16S sequences processed with Emu.
Collaborators
- Alona Tyshaieva (Univ of Dusseldorf)
- Dr. Alex Dilthey (Univ of Dusseldorf)
- Dr. Sonia Villapol (Houston Methodist)
- Dr. Tor Savidge (Texas Childrens)
- Dr. Qinglong Wu (Texas Childrens)
